In the late 1940s, Dr. Conrad Waddington, an embryologist, coined the term epigenetics. At that time, most embryologists did not consider genes important for human development. They believed genes only played a minor role, such as with eye color and hair color.
Dr. Waddington proposed that genes were regulated by an epigenetic landscape, and this landscape was to “illustrate the various developmental pathways a cell might take toward differentiation.”1 Scientists are just now beginning to understand how the epigenetic landscape that Dr. Waddington proposed more than 50 years ago applies to disease development and possible treatment.
The most commonly discussed disease associated with the study of epigenetics is cancer. An example of a discovery using epigenetics in cancer is the mutations in the BRCA1 gene linked to familial breast cancer. The BRCA1 gene acts as a tumor suppressor; scientists have discovered over 1,800 mutations associated with this gene. When there is a mutation in the BRCA1 gene, it can “trigger cells to grow and divide uncontrollably to form a tumor.”2
There seems to be a lot to still learn about epigenetics. Some studies have shown different polymorphisms and mutations that are associated with oral health, and many have the potential to help us better understand the development, progression, and susceptibility of dental disease.
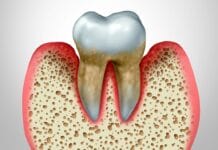
Periodontal Disease
According to the Centers for Disease Control and Prevention (CDC), 47% of adults have some form of periodontal disease. Compare that to 9.4% of Americans who have diabetes, and you can see that periodontal disease is one of the most common chronic diseases.3,4
Epigenetic modifications that contribute to periodontal disease can be attributed to environmental factors and inherited genome.5,6 Epigenetics is where these environmental factors and inherited genome interact, leading to susceptibility of the host to develop periodontal disease. The infection caused by oral pathogens, such as P. gingivalis and F. nucleatum, has the potential to induce changes in the epigenome, contributing to host susceptibility.6 The mechanism by which these changes occur include DNA methylation and histone modification.5
DNA methylation is when a methyl group is added to the DNA. This often modifies the function of the gene and affects gene expression as well as inhibits transcription of the gene. In periodontal disease, modification of the gene occurs locally where the biofilm and gingiva meet. This can cause one site to be periodontally involved, while another site in the same individual is healthy.5 DNA methylation contributes to gene silencing, which ultimately alters cytokine levels and influences immune response.7 These genes that alter the body’s immune response lend to the reason periodontal disease is a chronic disease.
Several different modifications are associated with histones. Acetylation and deacetylation are the two modifications found to increase susceptibility to periodontal disease. Histones are a group of proteins found in chromatin. Histone proteins assist in the transcription process. Their modification can either allow gene expression (acetylation) or suppress gene expression (deacetylation). This process also regulates cytokines and has been shown to be a key factor in inflammatory diseases.8
The good news is that epigenetic mechanisms are reversible, which makes them a great target in treating periodontal disease. Epigenetic drugs have been available since the late 1960s to treat cancer, though only within the last 20 years was the mechanism of action understood.9 Research in the relationship between epigenetics and periodontal disease is still in the early stages. This research lends hope, though, to the possibility of better treatment and prognosis for periodontal disease.
Pulp Inflammation
The most important etiological factor in pulp inflammation is bacteria. Pulp inflammation is associated with an accumulation of inflammatory mediators, cytokines, and chemokines. These inflammatory mediators are stimulated by bacteria toxins and are meant to assist in the reparative process. Because epigenetic modifications are believed to be reversible, better understanding of these changes may lead to new therapies for pulp inflammation.
Studies show inflamed pulp tissue affects epigenetic modifications; these modifications might affect dental pulp cells’ response to harmful stimuli.10 The interferon-gamma gene, a dimerized soluble cytokine, was found to be unmethylated or partially methylated in 93% of inflamed pulp tissue samples. Pulp tissue with complete methylation had no interferon-gamma gene transcription.
Pulp inflammation is one of the most difficult diagnoses to make in dentistry. These studies that have observed differing methylation patterns in healthy pulp tissue and inflamed pulp tissue may be the beginning of more accurate diagnosis of pulp inflammation.
In addition, the potential for developing therapeutic treatments for pulp inflammation using non-coding RNA has become a topic that is being investigated. Non-coding RNA is associated with odontoblast differentiation and may be the perfect therapeutic treatment in dentine formation and pulp mineralization.
Although this is very promising, there are a few concerns, including the possibility of neoplastic transformation and tumorigenesis. This is because epigenetic modification plays a role in cell development and differentiation. Further studies are needed to better understand epigenetics role in pulp inflammation and the development of effective and safe therapeutic treatments.11
Oral Squamous Cell Carcinoma
Oral squamous cell carcinomas (OSCC) are the most common oral malignancies. The development of oral cancer is due to a combination of genetic factors, as well as several environmental risk factors, including tobacco use, alcohol consumption, chronic inflammation, and viral infections, e.g., HPV. These factors can lead to alterations in oncogenes and tumor suppressor genes.
Studies show the epigenetic landscape is different between malignant and normal cells. This influences the development and progression of malignancies. As mentioned previously, the fact that these modifications are reversible opens new concepts for impairing cancer development.12
A recently discovered class of micro ribonucleic acid (miRNA) appears to play a crucial role in gene expression and the progression of malignancies. These studies indicate miRNA works as a tumor suppressor; however, it may undergo epigenetic silencing in the presence of malignancies, allowing oncogenes to proliferate, leading to the progression of malignancies.13
Studies have also shown that analysis of specific gene hypermethylation indicates that epigenetic status can also be used as a tool to predict prognosis. Epigenetic modifications in the structure of DNA can be inherited. These modifications can be preserved for multiple generations, contributing to inherited increased risk of OSCC.12
The study of epigenetics as it relates to OSCC has opened a door to better treatment. By combining standard chemotherapy with epigenetic modulatory drugs, we could see improvement in clinical outcomes.
Chemo resistant genes have been shown to be hypermethylated. The use of an epigenetic modulating drug can contribute to restoring the genes’ initial identity and reducing pathogenicity. Natural products are also being explored for use in epigenetic modification. In vitro green tea was administered to oral cancer models, and inhibitory effects that were related to a reversal of gene hypermethylation was observed.12
Although research in the area of epigenetics is still emerging, these findings are very exciting. I hope to see changes in treatment options for oral cancer and periodontal disease, as well as better diagnostic approaches for pulp inflammation.
This is just the tip of the iceberg. Epigenetics has been a game-changer for medicine, and I hope it will trickle into dentistry as well.
Now Listen to the Today’s RDH Dental Hygiene Podcast Below:
References
- Navis, A.R. Epigenetic Landscape. The Embryo Project Encyclopedia. 2007-10-30. Retrieved from https://embryo.asu.edu/pages/epigenetic-landscape
- U.S. National Library of Medicine. Genetics Home Reference. BRCA1 gene. Retrieved from https://ghr.nlm.nih.gov/gene/BRCA1#conditions
- Centers for Disease Control and Prevention. Periodontal Disease. Retrieved from https://www.cdc.gov/oralhealth/conditions/periodontal-disease.html
- Centers for Disease Control and Prevention. New CDC Report: More than 100 million Americans have diabetes or prediabetes. Retrieved from https://www.cdc.gov/media/releases/2017/p0718-diabetes-report.html
- Ari, G., Cheruki, S., Namasivayam, A. Epigenetics and Periodontitis: A Contemporary Review. J Clin Diagn Res. 2016 Nov; 10(11): ZE07-ZE09. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5198474/
- Larsson, L. Current Concepts of Epigenetics and its Role in Periodontitis. Curr Oral Health Rep. 2017; 4(4): 286-293. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5688199/
- Lavu, V., Venkatesan, V., Rao, S.R. The Epigenetic Paradigm in Periodontal Pathogenesis. J Indian Soc Periodontol. 2015 Mar-Apr; 19(2): 142-149. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4439621/
- Chaurasia, A. Epigenetics in Periodontal Diseases. J Clin Epigenet. 2017; 3(3):30. Retrieved from http://clinical-epigenetics.imedpub.com/epigenetics-in-periodontal-diseases.pdf
- Heerboth, S., Lapinska, K., Snyder, N., Leary, M., Rollinson, S., Sarkar, S. Use of Epigenetic Drugs in Disease: An Overview. Genet Epigenet. 2014; 6: 9-19. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4251063/
- Hui, T., Wang, C., Chen, D., Zheng, L., Huang, D., Ye, L. Epigenetic Regulation in Dental Pulp Inflammation. Oral Dis. 2017 Jan; 23(1): 22-28. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4993683/
- Kearney, M., Cooper, P.R., Smith, A.J., Duncan, H.F. Epigenetic Approaches to the Treatment of Dental Pulp Inflammation and Repair: Opportunities and Obstacles. Front Genet. 2018; 9: 311. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6090030/
- Irimie, A.I., Ciocan, C., Gulei, D., Mehterov, N., Atanasov, A.G., Dudea, D., Berindan-Neagoe, I. Current Insights into Oral Cancer Epigenetics. Int Mol Sci. 2018 Mar; 19(3): 670. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5877531/
- Hema, K.N., Smitha, T., Sheethal, H.S., Mirnalini, A. Epigenetics in Oral Squamous Cell Carcinoma. J Oral Maxillofac Pathol. 2017 May-Aug; 21(2): 252-259. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5596676/